Prokaryotic Escherichia coli protein expression
Prokaryotic Escherichia coli protein expression
 Yeast protein expression
Yeast protein expression
 Protein expression of baculovirus - insect cell
Protein expression of baculovirus - insect cell
 Protein expression in mammalian cells
Protein expression in mammalian cells
 Proteins are purified in small quantities and large quantities
Proteins are purified in small quantities and large quantities
 E. coli and yeast fermentation
E. coli and yeast fermentation
 Construction of cDNA library, yeast double hybridization, GST pulldown, Coip and other studies on pr
Construction of cDNA library, yeast double hybridization, GST pulldown, Coip and other studies on pr
 Gene cloning, gene mutation, and gene knockout
Gene cloning, gene mutation, and gene knockout

The yeast expression system has the advantages of both prokaryotic and advanced eukaryotic systems. It has the characteristics of common culture conditions, fast growth rate, relatively low cost, the ability to process proteins after translation, and easy access to soluble active recombinant proteins, etc., and is widely used in the production and preparation of recombinant proteins, especially eukaryotic proteins. Among them, Pichia pastoris has become a priority for yeast to express recombinant proteins due to its advantages of fast growth, abundant commercial expression vectors and high efficient secretion and expression.
1. Different vectors, different affinity tags (HIS, FC), secretion or intracellular expression can be selected after 1 to 2 weeks of shuttle vector construction; Our company can provide PPIC9K and PPICZAA secreted expression vectors free of charge. At the same time, we can provide our company's self-constructed PPIC9K-SUMO (mutation) vector to customers according to customer requirements and special conditions such as small target protein (less than 8KD). In this way, the expression level can be greatly improved and the characteristics of small protein are easy to degrade and not easy to purify can be overcome (1500 yuan is needed).
2.According to customer requirements and gene characteristics, suitable endonuclease linearized recombinant plasmid was selected and electrically transformed into Pichia pastoris GS115 or X-33, and the expected methanol utilization phenotype of the transformant was obtained.
3. Screening of high-copy yeast inverters (optional) for 2-3 weeks. If the GS115/ PPIC9K system was selected, G418 resistance screening (EG :0.5mg/mL, 1mg/mL, 2mg/mL) was performed after the first round of nutritional deficiency screening. If the X33/ PPICZAA system is selected, this step is combined with Step 2 in a gradient pressure screening process;
4 PCR identification of yeast positive transformants Several strains of high-copy yeast transformants were selected in 1 week, and the corresponding genomic DNA was extracted for identification by PCR (primers 5AOX1 and 3AOX1 could simultaneously identify the methanol utilization phenotype of the transformants).
5. Induced expression and isolation and purification of recombinant protein A few strains of high-copy yeast transformers (5-10 strains) were selected for small amount of induction expression (methanol as inducer) in one week, and the yeast induced superintents were collected at different time points for SDS-PAGE electrophoresis or DOT-blot analysis, so as to determine the optimal sampling time. Yeast inverters with the highest expression level (the same step 3 can be mutually verified);
6. Separation and purification of recombinant proteins for 1 week. The suitable chromatographic methods were selected to purify recombinant proteins. The commonly used chromatographic methods were: affinity chromatography (Ni, Proteina), anion and cation exchange chromatography (Q, SP), molecular exclusion chromatography, etc.
The purity of the recombinant protein was detected by SDS-PAGE electrophoresis, the concentration of the recombinant protein was quantified by BCA, and the immunogenicity of the recombinant protein was detected by Western-blot.
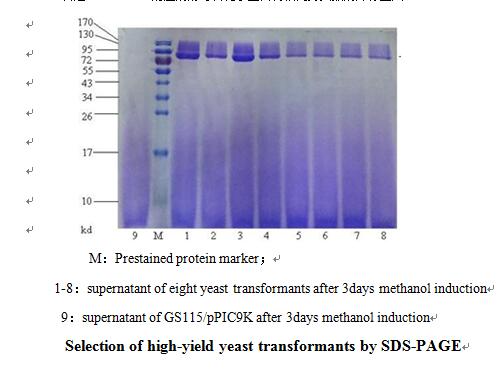
Case study: Secretory expression of a recombinant enzyme in Pichia pastoris (GS115/ PPIC9K system)
Background: The recombinant enzyme gene was inserted into PPIC9K vector by seamless cloning technology, and then electrotransformed into Pichia pastoris GS115, coated with MD plate for preliminary screening, and then several yeast inverters (300 strains of >) on MD plate were selected for G418 gradient screening (G418 concentration was set as 0.5mg/ml, 1mg/ml, 2mg/ml gradient). Finally, 8 strains of yeast transformants were selected from plates containing different concentrations of G418 and expressed with methanol. After 3 days, the expression superintents of each transformant were harvested for SDS-PAGE detection.
Conclusion: It can be seen from the figure that the expression levels of the 8 yeast transformants are ranked as follows: translane 1,3> track,4> track 5,6,7,8. According to the horizontal comparison of expression levels and combined with the gray scanning results of BandScan software, it is highly possible that: Trans 1 and 3 had the same copy number of foreign genes, Trans 2 and 4 had the same copy number of foreign genes, Trans 5, 6, 7 and 8 had the same copy number of foreign genes.